Fluoroscopy Simulator#
GPU-accelerated fluoroscopy (X-ray) simulation from CT volumes using differentiable ray marching.
Overview#
The fluorosim package generates realistic simulated X-ray images from CT volumes using Beer-Lambert physics and GPU-accelerated rendering via NVIDIA Slang with automatic differentiation.
Key Capabilities:
- Generate Digitally Reconstructed Radiographs (DRRs) from CT volumes at arbitrary C-arm poses
- Compute exact gradients for 2D/3D registration via Slang's compiler-level autodiff
- Achieve real-time performance (~150+ FPS on RTX A6000)
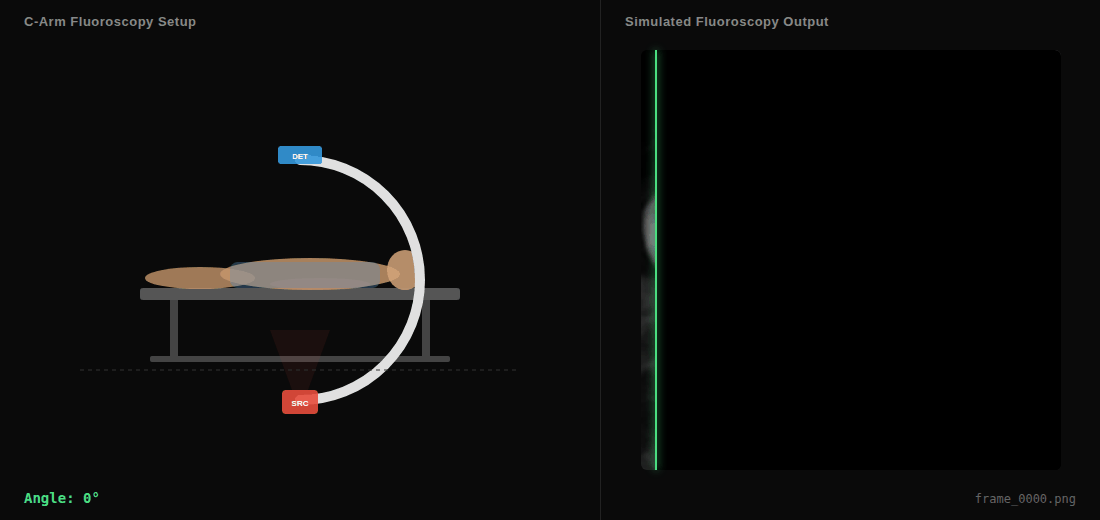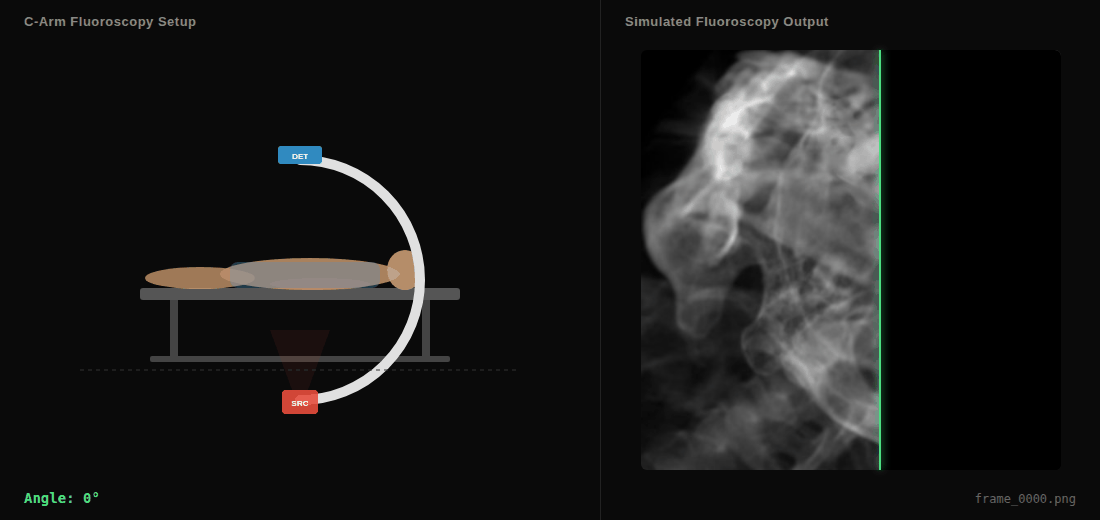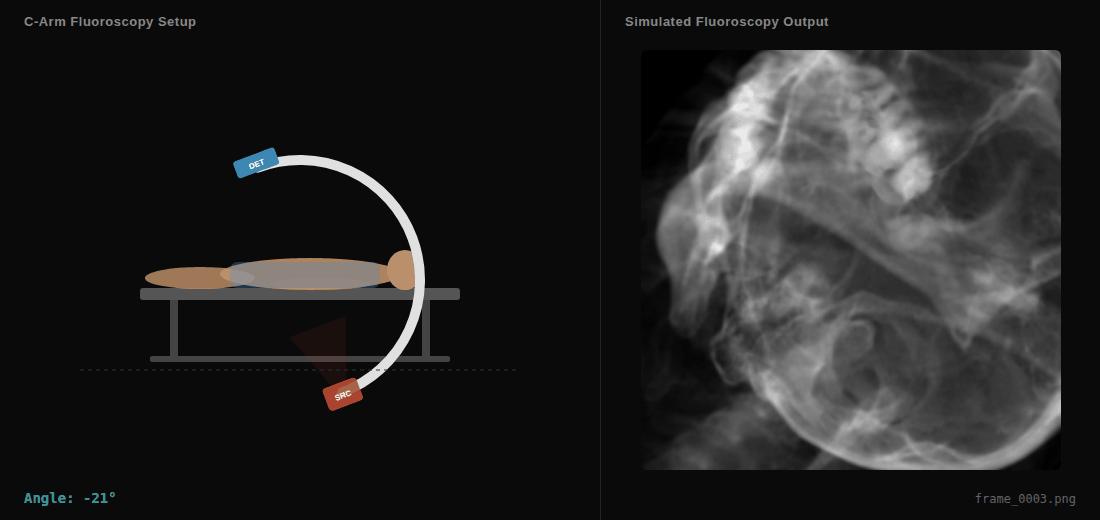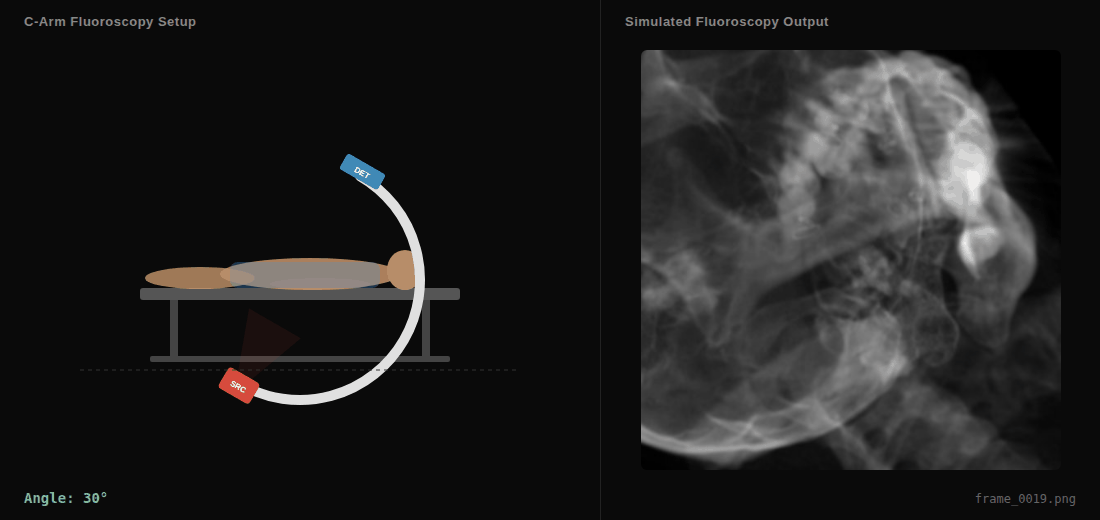
C-arm sweep animation showing the virtual X-ray source (SRC), detector (DET), and simulated fluoroscopy output in real-time.
Table of Contents#
- Installation
- Quick Start
- Architecture, API & Configuration — rendering pipeline, physics, API reference, C-arm configuration
- Examples & Test Data — test datasets, example scripts, running the examples
Installation#
Option 1: Using the I4H CLI (Recommended)#
The ./i4h CLI builds and runs inside a Docker container with all dependencies pre-installed.
Each step is a separate mode; the cached preprocessed volume persists between runs in fluoro-simulator/output/.
# Synthetic data workflow (no real CT data needed)
./i4h run fluoro-simulator preprocess_synthetic
./i4h run fluoro-simulator demo
# Real CT data workflow
./i4h run fluoro-simulator download_data
./i4h run fluoro-simulator preprocess_dicom
./i4h run fluoro-simulator demo
# List available modes
./i4h modes fluoro-simulator
# Launch an interactive shell inside the container
./i4h run-container fluoro-simulator
Set FLUOROSIM_OUTPUT_DIR or FLUOROSIM_CACHE_DIR to override the default output paths.
Option 2: Docker#
cd fluoro-simulator
# Build the Docker image
docker build -t fluorosim .
# Run the container
docker run -it --rm --gpus all fluorosim bash
# Inside the container
# Run the synthetic demo
python examples/preprocess_ct.py --synthetic
python examples/fluorosim_demo.py
# Run the demo with real CT data
# Download the dataset
kaggle datasets download -d adamhuan/multiphase-ct-anigography-2-datasets
unzip multiphase-ct-anigography-2-datasets.zip
python examples/preprocess_ct.py --dicom excellent/excellent/0
python examples/fluorosim_demo.py
Option 3: Bare Metal Installation#
cd fluoro-simulator
pip install -e .[all]
Requirements:
| Dependency | Purpose |
|---|---|
| CUDA-capable GPU | GPU-accelerated rendering |
slangpy >= 0.40 |
Slang shader compilation and autodiff |
numpy |
Array operations |
SimpleITK |
DICOM/NIfTI loading (optional) |
torch |
PyTorch integration for autograd (optional) |
Quick Start#
from fluorosim import VolumePreprocessor, FluoroSimulator, SimulatorConfig, PreprocessedVolume
# Step 1: Preprocess CT volume (HU → μ conversion)
volume = VolumePreprocessor.from_nifti("ct.nii.gz").preprocess()
# Step 2: Create simulator and render
simulator = FluoroSimulator(volume)
frame = simulator.render_frame(rotation=(0, 0.5, 0)) # 0.5 rad Y rotation
# Step 3: Save result
frame.save("output.png")
Step-by-Step Explanation#
Step 1: Volume Preprocessing#
CT volumes store tissue density in Hounsfield Units (HU). The VolumePreprocessor converts these to linear attenuation coefficients (μ in mm⁻¹) using a linear mapping:
μ = μ_min + (HU - HU_min) / (HU_max - HU_min) × (μ_max - μ_min)
Default mapping: HU ∈ [-1000, 3000] → μ ∈ [0.0, 0.02] mm⁻¹
# Load from various sources
volume = VolumePreprocessor.from_dicom("/path/to/dicom/").preprocess()
volume = VolumePreprocessor.from_nifti("ct.nii.gz").preprocess()
volume = VolumePreprocessor.from_numpy(hu_array, spacing_zyx_mm=(1.0, 0.5, 0.5)).preprocess()
# Cache to disk for fast reloading
volume = preprocessor.preprocess(output_dir="/tmp/fluoro_cache")
# Reload cached volume
volume = PreprocessedVolume.load("/tmp/fluoro_cache")
Step 2: Simulator Initialization#
The FluoroSimulator initializes the GPU rendering pipeline:
simulator = FluoroSimulator(volume, config=SimulatorConfig())
- uploads μ-volume to GPU as a 3D texture with trilinear interpolation
- compiles Slang shader with autodiff enabled
- configures virtual C-arm geometry (source, detector, isocenter)
Step 3: Frame Rendering#
Render a single X-ray frame at a specified C-arm pose:
frame = simulator.render_frame(
rotation=(rx, ry, rz), # Euler angles in radians
translation=(tx, ty, tz), # Translation in mm
)
| Parameter | Axis | Clinical Term | Example |
|---|---|---|---|
rx |
X | Cranial (+) / Caudal (−) | rx=0.1 → 5.7° cranial tilt |
ry |
Y | LAO (+) / RAO (−) | ry=0.5 → 28.6° LAO |
rz |
Z | Roll | rz=0 → no roll |
Step 4: Output#
# Access rendered image as numpy array
image = frame.image # Shape: (H, W), dtype: float32, range: [0, 1]
# Save to disk
frame.save("output.png") # 8-bit grayscale PNG
frame.save("output.npy") # Full-precision numpy
Rendering Multiple Frames (Cine Sequence)#
from fluorosim import Pose
import math
# Generate a LAO/RAO sweep animation
poses = [
Pose(rotation=(0, math.radians(angle), 0))
for angle in range(-30, 31, 2) # -30° to +30° in 2° steps
]
cine = simulator.render_cine(poses, fps=15.0)
cine.save_all("/tmp/output", format="png") # frame_0000.png, frame_0001.png, ...
# Access as numpy array
frames_array = cine.to_numpy() # Shape: (N, H, W)
Further Reading#
- Architecture, API & Configuration — Physics model, differentiable rendering, full API reference, and C-arm geometry configuration.
- Examples & Test Data — Recommended datasets (e.g. Kaggle), command-line examples, single-frame/cine/streaming code, and step-by-step instructions for
preprocess_ct.pyandfluorosim_demo.py.